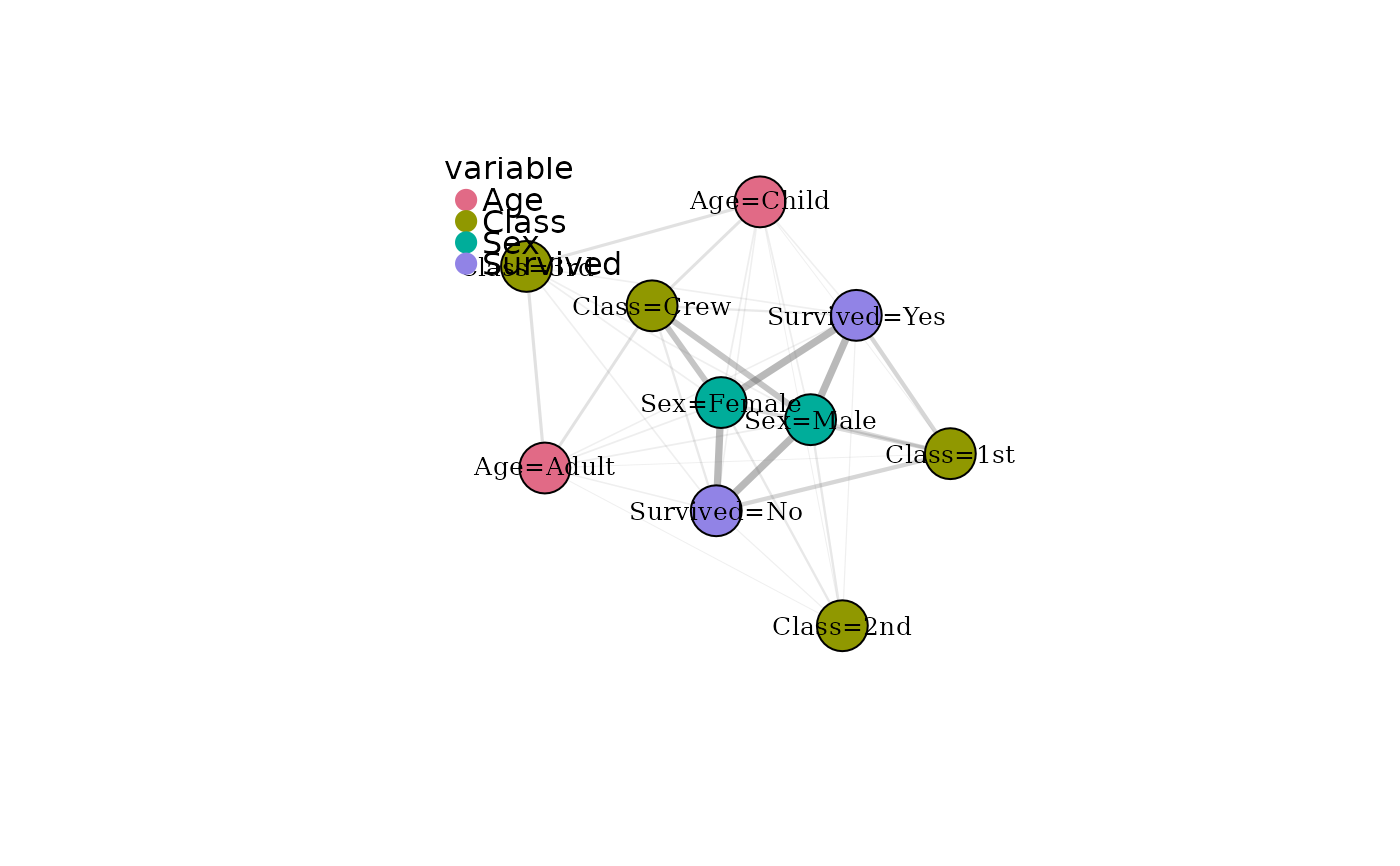
Visualises a catmodgraph object. Nodes represent modalities and
edges represent cross-variable modality associations. By default, node
colours indicate the originating variable. If modality communities have
been detected, colours can instead reflect community membership.
Arguments
- x
A
catmodgraphobject.- color_by
Character. One of
"variable"(default) or"cluster".- signed
Logical. If
TRUE, edge colour encodes the sign of the stored standardised Pearson residual: green for positive (attraction), red for negative (repulsion). Default isFALSE, which uses a uniform grey edge colour. Requires thestd_residedge attribute (present by default in graphs built withbuild_modality_graph).- show_labels
Logical. If
TRUE, node labels are drawn. Default isTRUE.- layout
Character. Graph layout passed to
igraph::layout_with_fr()origraph::layout_with_kk(). One of"fr"(default) or"kk".- vertex_size
Numeric. Node size. Default is
24.- edge_scale
Numeric. Multiplicative factor applied to edge widths. Default is
8.- remove_isolates
Logical. If
TRUE(default), vertices with degree 0 are hidden from the plot. Isolated modalities typically arise after pruning and carry no community structure information, so removing them improves interpretability. Set toFALSEto see every vertex in the graph. Does not modify the input object.- ...
Further arguments passed to
plot.igraph().
Details
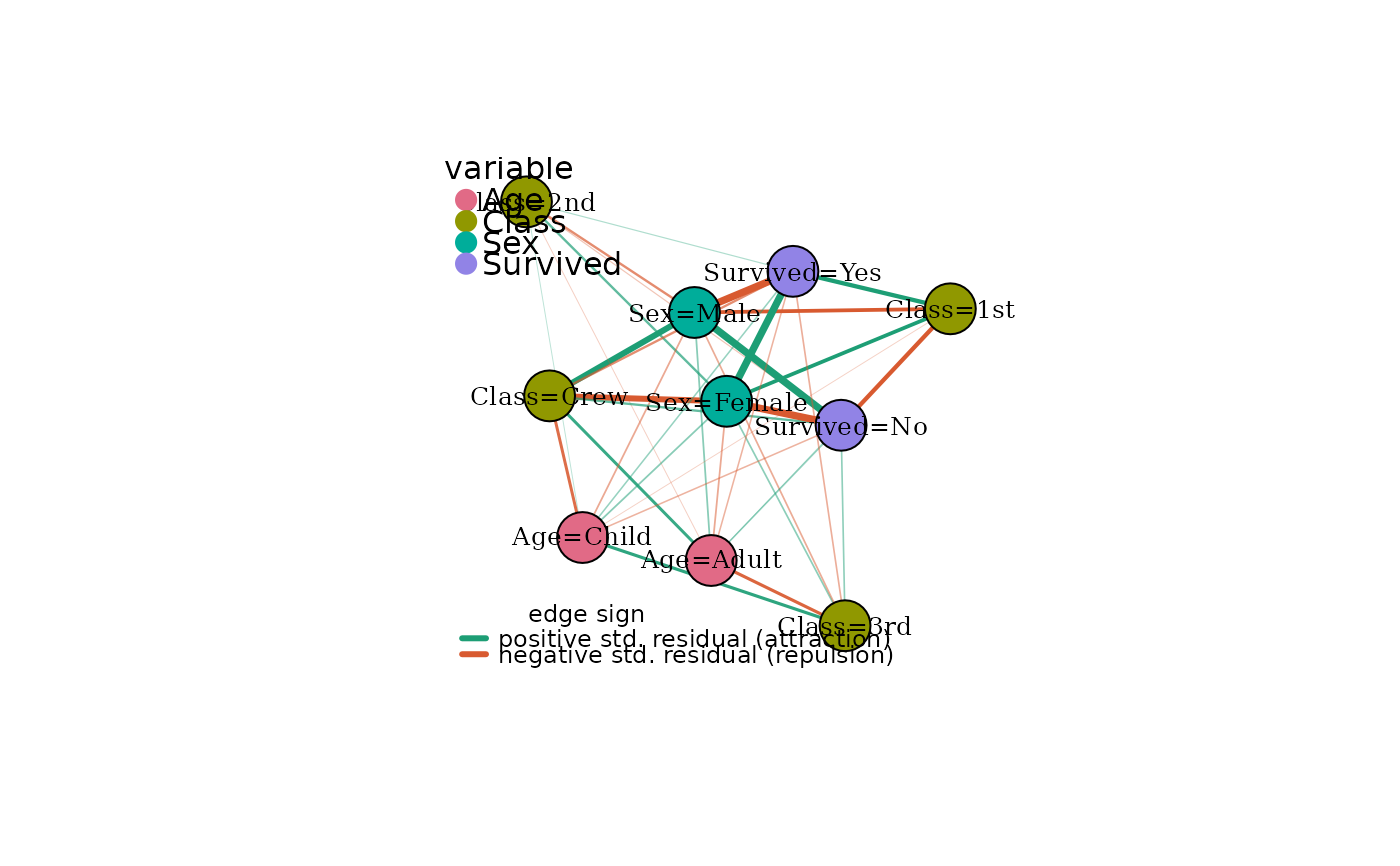
When signed = TRUE, edges are coloured by the sign of the stored
standardised Pearson residual (std_resid edge attribute): green
for positive (modalities co-occurring more than expected under
independence) and red for negative (co-occurring less than expected).
Edge alpha transparency then scales with |std_resid|.
Examples
df <- expand_table(Titanic)
mg <- build_modality_graph(df)
mg <- cluster_modalities(mg)
# Base plotting
plot(mg)
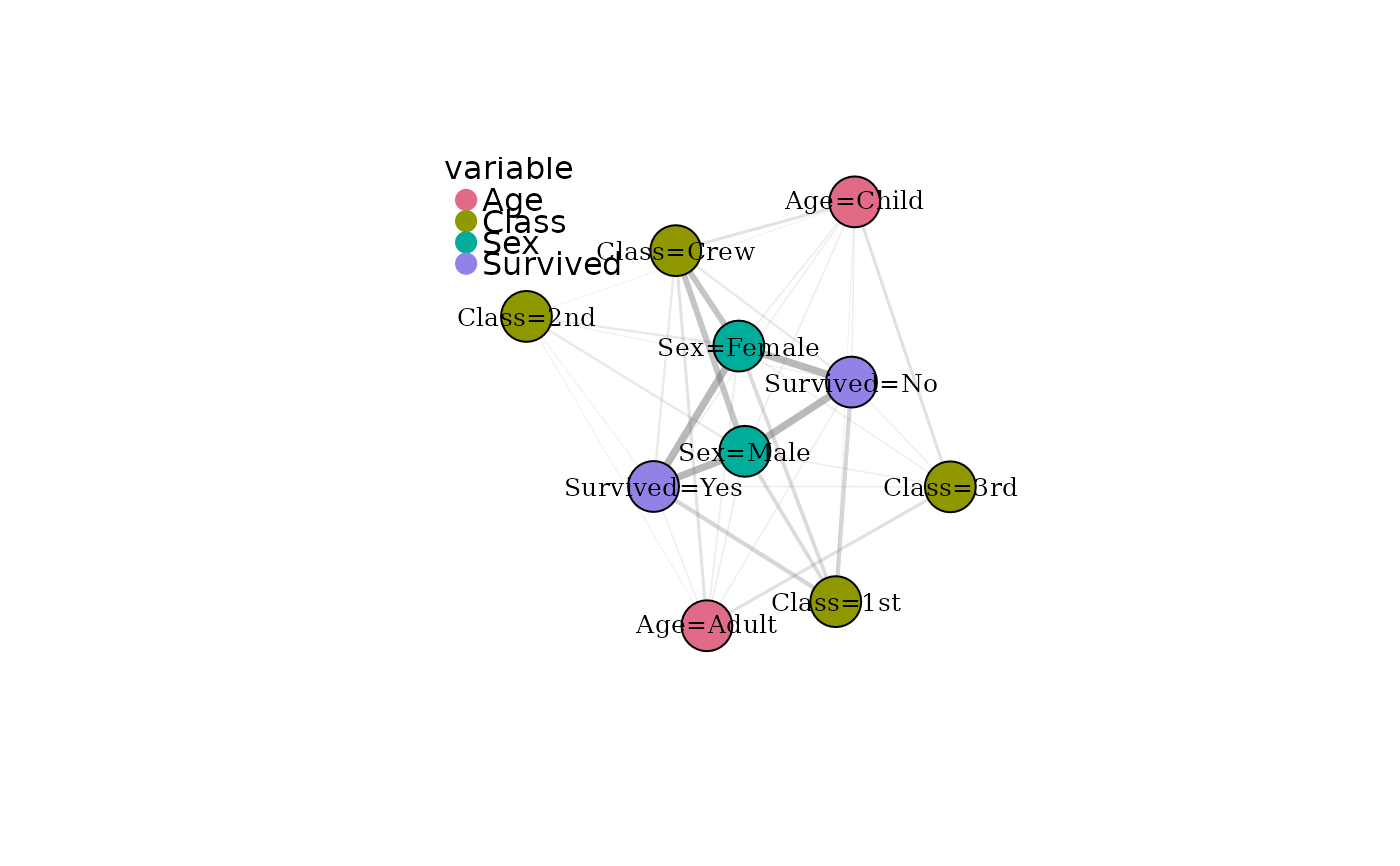 # Colour nodes by detected modality community
plot(mg, color_by = "cluster")
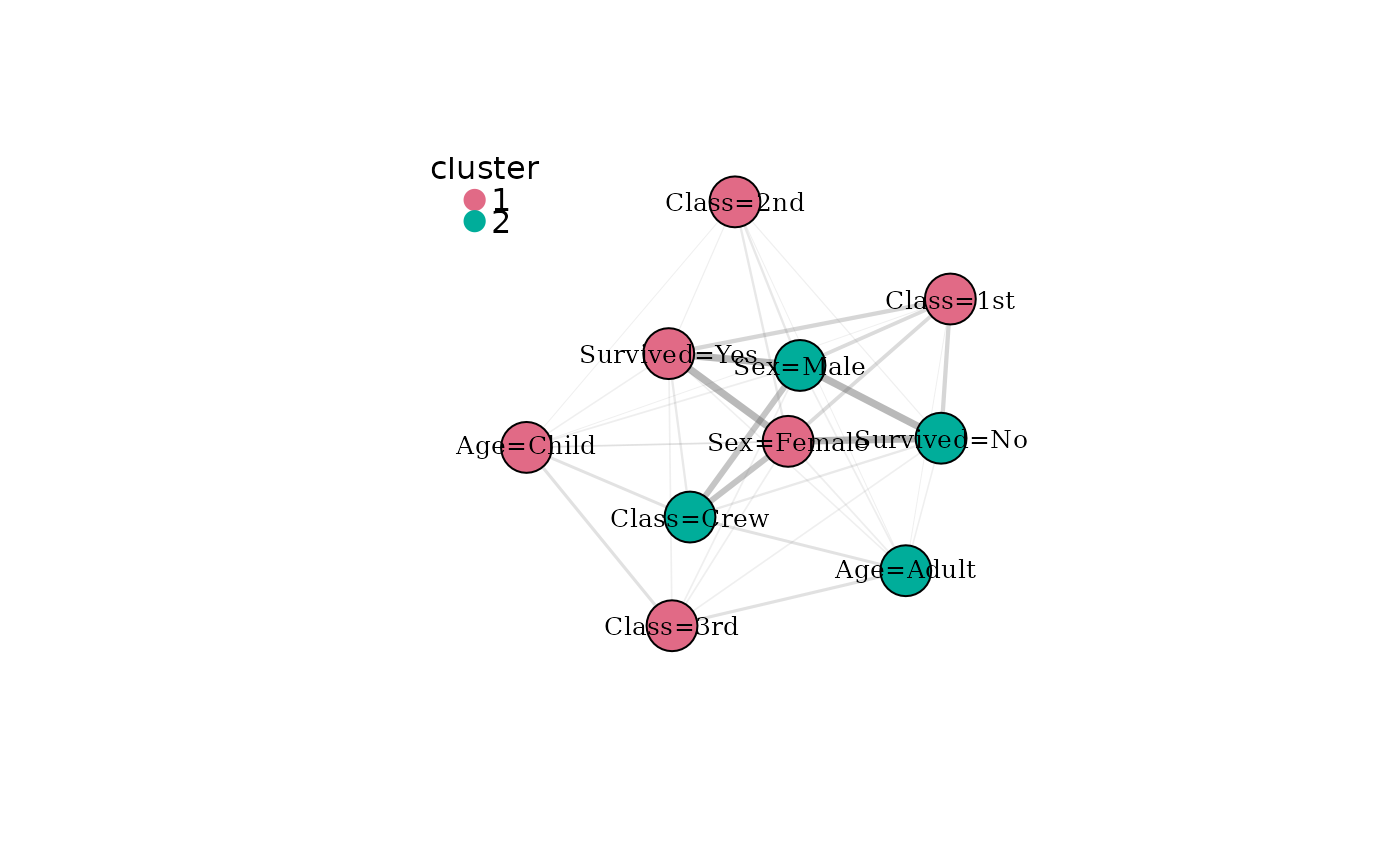
# Colour nodes by detected modality community
plot(mg, color_by = "cluster")
 # Signed edges: green = attraction, red = repulsion
plot(mg, signed = TRUE)
# Signed edges: green = attraction, red = repulsion
plot(mg, signed = TRUE)
 df <- expand_table(Titanic)
mg <- build_modality_graph(df)
mg <- cluster_modalities(mg)
# Base plotting
plot(mg)
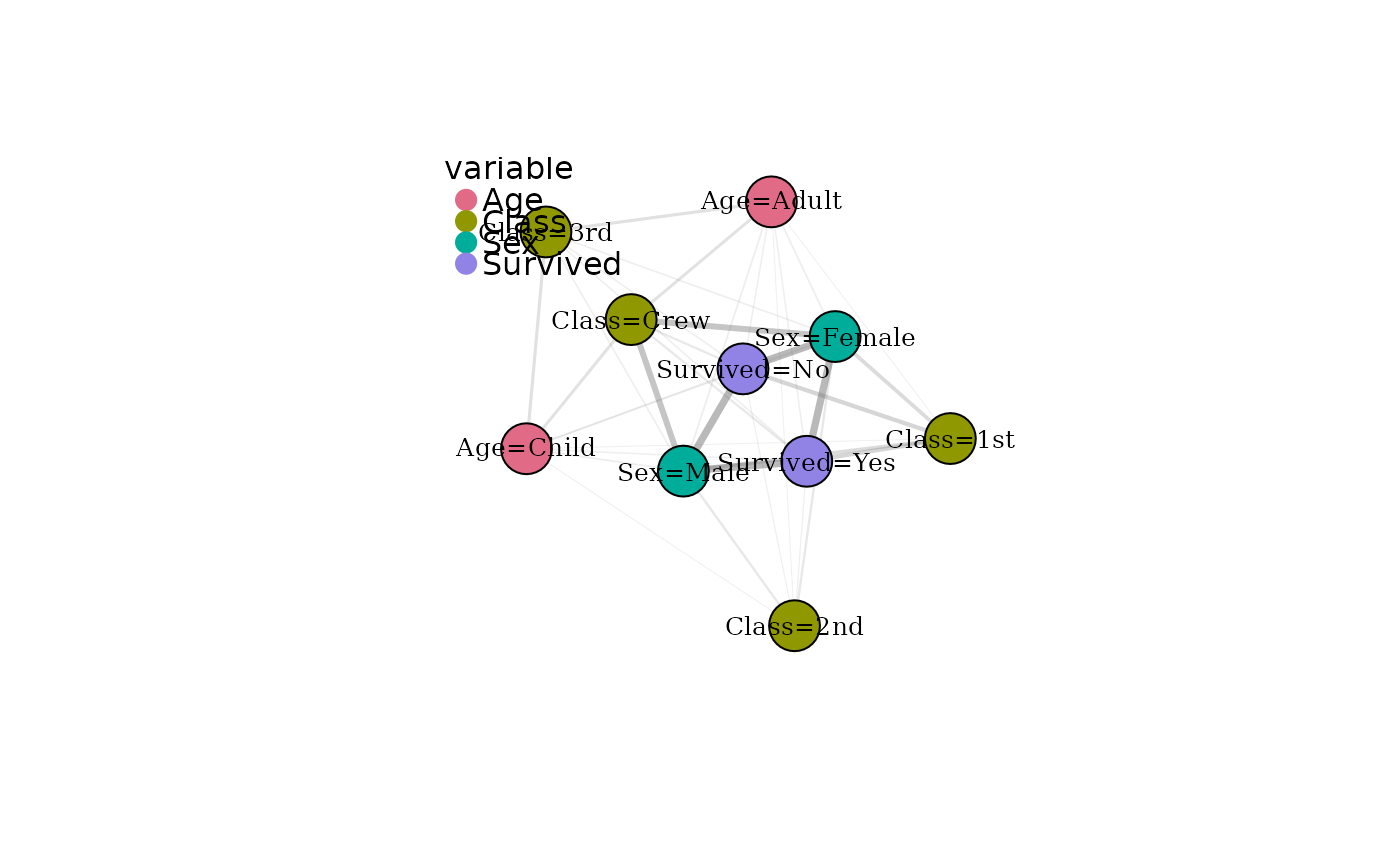
df <- expand_table(Titanic)
mg <- build_modality_graph(df)
mg <- cluster_modalities(mg)
# Base plotting
plot(mg)
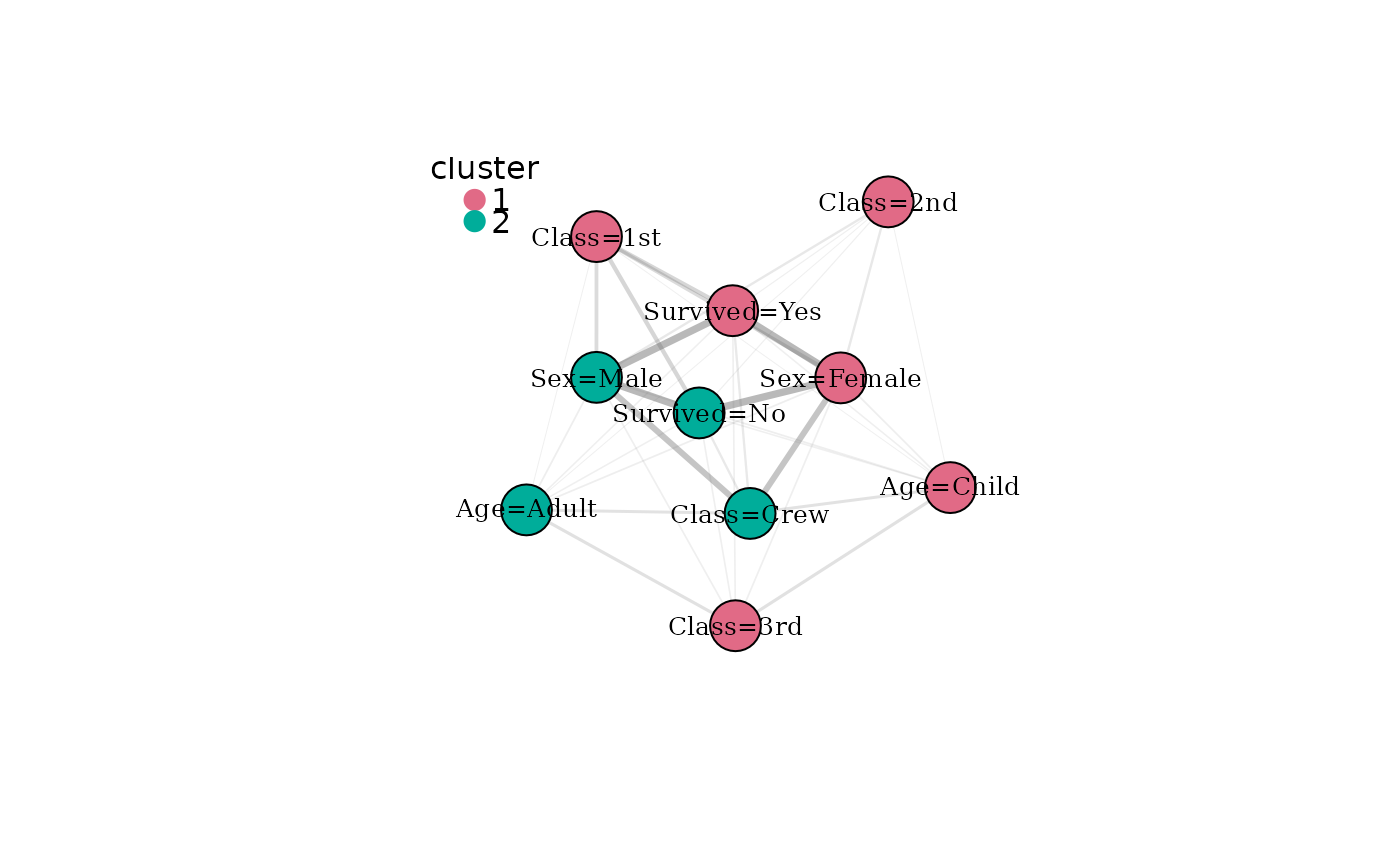
 # Colour nodes by detected modality community
plot(mg, color_by = "cluster")
# Colour nodes by detected modality community
plot(mg, color_by = "cluster")
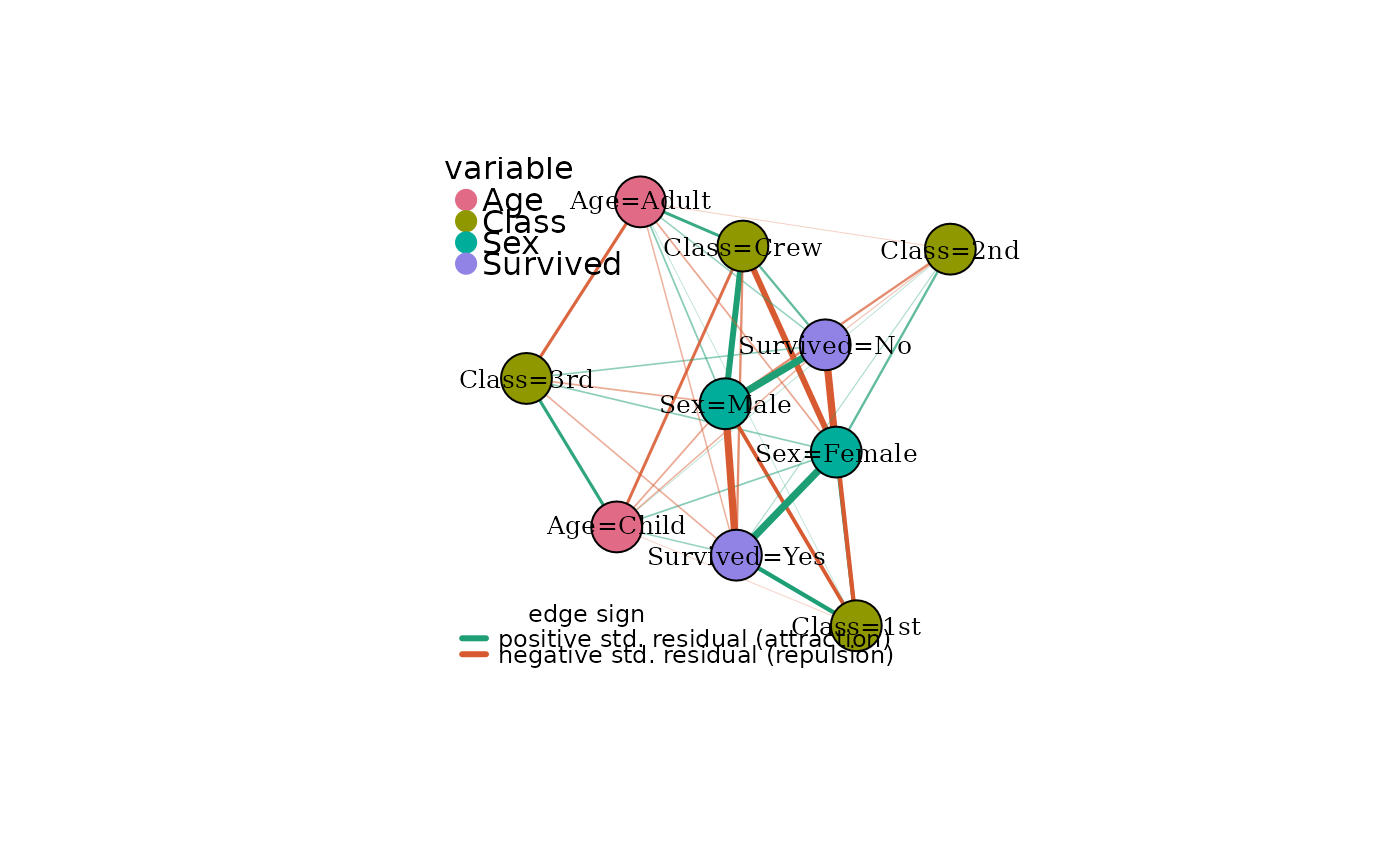
 # Signed edges: green = attraction, red = repulsion
plot(mg, signed = TRUE)
# Signed edges: green = attraction, red = repulsion
plot(mg, signed = TRUE)
 # Show isolated vertices (not hidden by default)
plot(mg, remove_isolates = FALSE)
# Show isolated vertices (not hidden by default)
plot(mg, remove_isolates = FALSE)